Comparing Text using Soundex, PHONIC_ENCODE and FUZZY_MATCH
When comparing text strings we have a number of functions on Oracle Database to help us. These include SOUNDEX, PHONIC_ENCODE and FUZZY_MATCH. Let’s have a look at what each of these can do.
The SOUNDEX function returns a character string containing the phonetic representation of a string. SOUNDEX lets you compare words that are spelled differently, but sound alike in English. To illustrate this, let’s compare some commonly spelled words and their variations, for example McDonald, MacDonald, and Smith and Smyth.
select soundex('MCDONALD') from dual;
SOUNDEX('MCDONALD')
___________________
M235
SQL> select soundex('MACDONALD') from dual;
SOUNDEX('MACDONALD')
____________________
M235
SQL> pause next_smith_smyth
next_smith_smyth
SQL>
SQL> select soundex('SMITH') from dual;
SOUNDEX('SMITH')
________________
S530
SQL> select soundex('SMYTH') from dual;
SOUNDEX('SMYTH')
________________
S530
Here we get to see SOUNDEX function returns the same code for each of the spelling variations. This function can be easily used to search for and to compare text in our tables.
Now let’s have a look at some of the different ways to spell Brendan.
select soundex('Brendan'),
2 soundex('BRENDAN'),
3 soundex('Breandan'),
4 soundex('Brenden'),
5 soundex('Brandon'),
6 soundex('Brendawn'),
7 soundex('Bhenden'),
8 soundex('Brendin'),
9 soundex('Brendon'),
10 soundex('Beenden'),
11 soundex('Breenden'),
12 soundex('Brendin'),
13 soundex('Brendyn'),
14 soundex('Brandon'),
15 soundex('Brenainn'),
16 soundex('Bréanainn')
17 from dual;
SOUNDEX('BRENDAN') SOUNDEX('BRENDAN') SOUNDEX('BREANDAN') SOUNDEX('BRENDEN') SOUNDEX('BRANDON') SOUNDEX('BRENDAWN') SOUNDEX('BHENDEN') SOUNDEX('BRENDIN') SOUNDEX('BRENDON') SOUNDEX('BEENDEN') SOUNDEX('BREENDEN') SOUNDEX('BRENDIN') SOUNDEX('BRENDYN') SOUNDEX('BRANDON') SOUNDEX('BRENAINN') SOUNDEX('BRÉANAINN')
__________________ __________________ ___________________ __________________ __________________ ___________________ __________________ __________________ __________________ __________________ ___________________ __________________ __________________ __________________ ___________________ ____________________
B653 B653 B653 B653 B653 B653 B535 B653 B653 B535 B653 B653 B653 B653 B655 B655 We can see which variations of my name can be labeled as being similar in sound.
An alternative function is to use the PHONIC_ENCODE. The PHONIC_ENCODE function converts text into language-specific codes based on the pronunciation of the text. It implements the Double Metaphone algorithm and an alternative algorithm. DOUBLE_METAPHONE returns the primary code. DOUBLE_METAPHONE_ALT returns the alternative code if present. If the alternative code is not present, it returns the primary code.
select col, phonic_encode(double_metaphone,col) double_met
2 from ( values
3 ('Brendan'), ('BRENDAN'), ('Breandan'), ('Brenden'), ('Brandon'),
4 ('Brendawn'), ('Bhenden'), ('Brendin'), ('Brendon'), ('Beenden'),
5 ('Breenden'), ('Brendin'), ('Brendyn'), ('Brandon'), ('Brenainn'),
6 ('Bréanainn')
7 ) names (col);
COL DOUBLE_MET
_________ __________
Brendan PRNT
BRENDAN PRNT
Breandan PRNT
Brenden PRNT
Brandon PRNT
Brendawn PRNT
Bhenden PNTN
Brendin PRNT
Brendon PRNT
Beenden PNTN
Breenden PRNT
Brendin PRNT
Brendyn PRNT
Brandon PRNT
Brenainn PRNN
Bréanainn PRNN The final function we’ll look at is FUZZY_MATCH. The FUZZY_MATCH function is language-neutral. It determines the similarity between two strings and supports several algorithms. Here is a simple example, again using variations of Brendan
with names (col) as (
2 values
3 ('Brendan'), ('BRENDAN'), ('Breandan'), ('Brenden'), ('Brandon'),
4 ('Brendawn'), ('Bhenden'), ('Brendin'), ('Brendon'), ('Beenden'),
5 ('Breenden'), ('Brendin'), ('Brendyn'), ('Brandon'), ('Brenainn'),
6 ('Bréanainn') )
7 select a.col, b.col, fuzzy_match(levenshtein, a.col, b.col) as lev
8 from names a, names b
9 where a.col != b.col;
COL COL LEV
________ _________ ___
Brendan BRENDAN 15
Brendan Breandan 88
Brendan Brenden 86
Brendan Brandon 72
Brendan Brendawn 88
Brendan Bhenden 72
Brendan Brendin 86
Brendan Brendon 86
Brendan Beenden 72
Brendan Breenden 75
Brendan Brendin 86
Brendan Brendyn 86
Brendan Brandon 72
Brendan Brenainn 63
Brendan Bréanainn 45
...Only a portion of the output is shown above. Similar solution would be the following and additionally we can compare the outputs from a number of the algorithms.
with names (col) as (
values
('Brendan'), ('BRENDAN'), ('Breandan'), ('Brenden'), ('Brandon'),
('Brendawn'), ('Bhenden'), ('Brendin'), ('Brendon'), ('Beenden'),
('Breenden'), ('Brendin'), ('Brendyn'), ('Brandon'), ('Brenainn'),
('Bréanainn') )
select a.col as col1, b.col as col2,
fuzzy_match(levenshtein, a.col, b.col) as lev,
fuzzy_match(jaro_winkler, a.col, b.col) as jaro_winkler,
fuzzy_match(bigram, a.col, b.col) as bigram,
fuzzy_match(trigram, a.col, b.col) as trigram,
fuzzy_match(whole_word_match, a.col, b.col) as jwhole_word,
fuzzy_match(longest_common_substring, a.col, b.col) as lcs
from names a, names b
where a.col != b.col
and ROWNUM <= 10;
COL1 COL2 LEV JARO_WINKLER BIGRAM TRIGRAM JWHOLE_WORD LCS
_______ ________ ___ ____________ ______ _______ ___________ ___
Brendan BRENDAN 15 48 0 0 0 14
Brendan Breandan 88 93 71 50 0 50
Brendan Brenden 86 94 66 60 0 71
Brendan Brandon 72 84 50 0 0 28
Brendan Brendawn 88 97 71 66 0 75
Brendan Bhenden 72 82 33 20 0 42
Brendan Brendin 86 94 66 60 0 71
Brendan Brendon 86 94 66 60 0 71
Brendan Beenden 72 82 33 20 0 42
Brendan Breenden 75 90 57 33 0 37 By default, the output is a percentage similarity, but the UNSCALED keyword can be added to return the raw value.
with names (col) as (
values
('Brendan'), ('BRENDAN'), ('Breandan'), ('Brenden'), ('Brandon'),
('Brendawn'), ('Bhenden'), ('Brendin'), ('Brendon'), ('Beenden'),
('Breenden'), ('Brendin'), ('Brendyn'), ('Brandon'), ('Brenainn'),
('Bréanainn') )
select a.col as col1, b.col as col2,
fuzzy_match(levenshtein, a.col, b.col, unscaled) as lev,
fuzzy_match(jaro_winkler, a.col, b.col, unscaled) as jaro_winkler,
fuzzy_match(bigram, a.col, b.col, unscaled) as bigram,
fuzzy_match(trigram, a.col, b.col, unscaled) as trigram,
fuzzy_match(whole_word_match, a.col, b.col, unscaled) as jwhole_word,
fuzzy_match(longest_common_substring, a.col, b.col, unscaled) as lcs
from names a, names b
where a.col != b.col
and ROWNUM <= 10;
COL1 COL2 LEV JARO_WINKLER BIGRAM TRIGRAM JWHOLE_WORD LCS
_______ ________ ___ ____________ ______ _______ ___________ ___
Brendan BRENDAN 6 0.48 0 0 0 1
Brendan Breandan 1 0.93 5 3 0 4
Brendan Brenden 1 0.94 4 3 0 5
Brendan Brandon 2 0.84 3 0 0 2
Brendan Brendawn 1 0.97 5 4 0 6
Brendan Bhenden 2 0.82 2 1 0 3
Brendan Brendin 1 0.94 4 3 0 5
Brendan Brendon 1 0.94 4 3 0 5
Brendan Beenden 2 0.82 2 1 0 3
Brendan Breenden 2 0.9 4 2 0 3This was another post in the ‘Back to Basics’ series of posts. Make sure to check out the other posts in the series.
SQL Firewall – Part 2
In a previous post, we’ve explored some of the core functionality of SQL Firewall in Oracle 23ai, In this post I’ll explore some of the other functionality that I’ve had to use as we’ve deployed SQL Firewall over the past few weeks.
Sometimes, when querying the DBA_SQL_FIREWALL_VIOLATIONS view, you might not get the current up to-date violations, or if you are running it for the first time you might get now rows or violations being returned from the view. This is a slight timing issue, as the violations log/cacbe might not have been persisted to the data dictionary. If you end up in this kind of situation you might need to flush the logs to to data dictionary. To do this, run the following.
exec dbms_sql_firewall.flush_logs;As you work with SQL Firewall on an ongoing basis, where you are turning it on and off at various stages, it can be easy to lose track of whether the Firewall is turned on or off. Being able to check the current status becomes important. To check the currect status, we can query DBA_SQL_FIREWALL_STATUS
select status
from dba_sql_firewall_status;
STATUS
——–
ENABLED
After checking that, we can then run either of the following.
exec dbms_sql_firewall.disable;or
exec dbms_sql_firewall.enableAfter creating your Allowed Lists for your various scenarios, at some point, you might need to add or remove individual statements/queries from a list. An important element for this is to locate the SQL_ID and SQL_SIGNATURE.
exec dbsm_sql_firewall.delete_allowed_sql(username => 'SCOTT', allowed_sql_id => 1);and to add a single statement
exec dbms_sql_firewall.append_allow_list_single_sql(username => 'SCOTT', sql_signature => '... ... ... ...',
current_user => 'PSMITH', top_level => 'Y',
source => DBMS_SQL_FIREWALL.VIOLATION_LOG);If you are using the Database Scheduler to run jobs, these will keep appearing in the Firewall logs. As these jobs run on a regular basis and new jobs can be added all the time, you will need to manage these. An alternative is to assume these jobs are safe to run. With SQL Firewall the managing of these can be very easy by getting SQL Firewall to ignore them.
exec dbms_sql_firewall.execlude(DBMS_SQL_FIREWALL.SCHEDULAR_JOB);SQL Firewall – Part 1
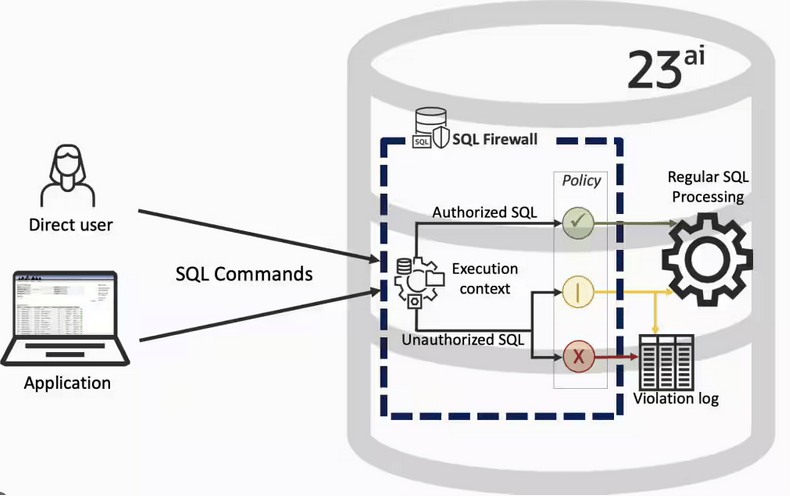
Typically, most IT architectures involve a firewall to act as a barrier, to monitor and to control network traffic. Its aim is to prevent unauthorised access and malicious activity. The firewall enforces rules to allow or block specific traffic (and commands/code). The firewall tries to protect our infrastructure and data. Over time, we have seen examples of how such firewalls have failed. We’ve also seen how our data (and databases) can be attacked internally. There are many ways to access the data and database without using the application. Many different people can have access to the data/database for many different purposes. There has been a growing need to push the idea of and the work of the firewall back to being closer to the data, that is, into the database.
SQL Firewall allows you to implement a firewall within the database to control what commands are allowed to be run on the data. With SQL Firewall you can:
- Monitor the SQL (and PL/SQL) activity to learn what the normal or typical SQL commands are being run on the data
- Captures all commands and logs them
- Manage a list of allowed commands, etc, using Policies
- Block and log all commands that are not allowed. Some commands might be allowed to run
Let’s walk through a simple example of setting this up and using it. For this example I’m assuming you have access to SYSTEM and another schema, for example SCOTT schema with the EMP, DEPT, etc tables.
Step 1 involves enabling SQL Firewall. To do this, we need to connect to the SYS schema and run the function to enable it.
grant sql_firewall_admin to system;Then connect to SYSTEM to enable the firewall.
exec dbms_sql_firewall.enable;For Step 2 we need to turn it on, as in we want to capture some of the commands being performed on the Database. We are using the SCOTT schema, so let’s capture what commands are run in that schema. [remember we are still connected to SYSTEM schema]
begin
dbms_sql_firewall.create_capture (
username=>'SCOTT',
top_level_only=>true);
end;
Now that SQL Firewall is running, Step 3, we can switch to and connect to the SCOTT schema. When logged into SCOTT we can run some SQL commands on our tables.
select * from dept;
select deptno, count(*) from emp group by deptno;
select * from emp where job = 'MANAGER';For Step 4, we can log back into SYSTEM and stop the capture of commands.
exec dbms_sql_firewall.stop_capture('SCOTT');We can then use the dictionary view DBA_SQL_FIREWALL_CAPTURE_LOG to see what commands were captured and logged.
column command_type format a12
column current_user format a15
column client_program format a45
column os_user format a10
column ip_address format a10
column sql_text format a30
select command_type,
current_user,
client_program,
os_user,
ip_address,
sql_text
from dba_sql_firewall_capture_logs
where username = 'SCOTT';The screen isn’t wide enough to display the results, but if you run the above command, you’ll see the three SELECT commands we ran above.
Other SQL Firewall dictionary views include DBA_SQL_FIREWALL_ALLOWED_IP_ADDR, DBA_SQL_FIREWALL_ALLOWED_OS_PROG, DBA_SQL_FIREWALL_ALLOWED_OS_USER and DBA_SQL_FIREWALL_ALLOWED_SQL.
For Step 5, we want to say that those commands are the only commands allowed in the SCOTT schema. We need to create an allowed list. Individual commands can be added, or if we want to add all the commands captured in our log, we can simple run
exec dbms_sql_firewall.generate_allow_list ('SCOTT');
exec dbms_sql_firewall.enable_allow_list (username=>'SCOTT',block=>true);Step 6 involves testing to see if the generated allowed list for SQL Firewall work. For this we need to log back into SCOTT schema, and run some commands. Let’s start with the three previously run commands. These should run without any problems or errors.
select * from dept;
select deptno, count(*) from emp group by deptno;
select * from emp where job = 'MANAGER';Now write a different query and see what is returned.
select count(*) from dept;
Error starting at line : 1 in command -
select count(*) from dept
*
ERROR at line 1:
ORA-47605: SQL Firewall violationOur new SQL command has been blocked. Which is what we wanted.
As an Administrator of the Database (DBA) you can monitor for violations of the Firewall. Log back into SYSTEM and run the following.
set lines 150
column occurred_at format a40
select sql_text,
firewall_action,
ip_address,
cause,
occurred_at
from dba_sql_firewall_violations
where username = 'SCOTT';
SQL_TEXT FIREWAL IP_ADDRESS CAUSE OCCURRED_AT
------------------------------ ------- ---------- ----------------- ----------------------------------------
SELECT COUNT (*) FROM DEPT Blocked 10.0.2.2 SQL violation 18-SEP-25 06.55.25.059913 PM +00:00
If you decide this command is ok to be run in the schema, you can add it to the allowed list.
exec dbms_sql_firewall.append_allow_list('SCOTT', dbms_sql_firewall.violation_log);The example above gives you the steps to get up and running with SQL Firewall. But there is lots more you can do with SQL Firewall, from monitoring of commands etc, to managing violations, to managing the logs, etc. Check out my other post covering some of these topics.
OCI Speech Real-time Capture
Capturing Speech-to-Text is a straight forward step. I’ve written previously about this, giving an example. But what if you want the code to constantly monitor for text input, giving a continuous. For this we need to use the asyncio python library. Using the OCI Speech-to-Text API in combination with asyncio we can monitor a microphone (speech input) on a continuous basis.
There are a few additional configuration settings needed, including configuring a speech-to-text listener. Here is an example of what is needed
lass MyListener(RealtimeSpeechClientListener):
def on_result(self, result):
if result["transcriptions"][0]["isFinal"]:
print(f"1-Received final results: {transcription}")
else:
print(f"2-{result['transcriptions'][0]['transcription']} \n")
def on_ack_message(self, ackmessage):
return super().on_ack_message(ackmessage)
def on_connect(self):
return super().on_connect()
def on_connect_message(self, connectmessage):
return super().on_connect_message(connectmessage)
def on_network_event(self, ackmessage):
return super().on_network_event(ackmessage)
def on_error(self, error_message):
return super().on_error(error_message)
def on_close(self, error_code, error_message):
print(f'\nOCI connection closing.')
async def start_realtime_session(customizations=[], compartment_id=None, region=None):
rt_client = RealtimeSpeechClient(
config=config,
realtime_speech_parameters=realtime_speech_parameters,
listener=MyListener(),
service_endpoint=realtime_speech_url,
signer=None, #authenticator(),
compartment_id=compartment_id,
)
asyncio.create_task(send_audio(rt_client))
if __name__ == "__main__":
asyncio.run(
start_realtime_session(
customizations=customization_ids,
compartment_id=COMPARTMENT_ID,
region=REGION_ID,
)
)Additional customizations can be added to the Listener, for example, what to do with the Audio captured, what to do with the text, how to mange the speech-to-text (there are lots of customizations)
Using Annotations to Improve Responses from SelectAI
In my previous posts on using SelectAI, I illustrated how adding metadata to your tables and columns can improve the SQL generated by the LLMs. Some of the results from those where a bit questionable. Going forward (from 23.9 onwards, although it might get backported), it appears that we need to add additional metadata to obtain better responses from the LLMs, by way of Annotations. Check out my previous posts on SelectAI at post-1, post-2, post-3, post-4, post-5.
Let’s have a look at how to add Annotations to support SelectAI with generating better responses.
NB: Support for additional LLMs is constantly being updated. Check out the current list here.
The following is an example from my previous post on adding table and column comments.
CREATE TABLE TABLE1(
c1 NUMBER(2) not null primary key,
c2 VARCHAR2(50) not null,
c3 VARCHAR2(50) not null);
COMMENT ON TABLE table1 IS 'Department table. Contains details of each Department including Department Number, Department Name and Location for the Department';
COMMENT ON COLUMN table1.c1 IS 'Department Number. Primary Key. Unique. Used to join to other tables';
COMMENT ON COLUMN table1.c1 IS 'Department Name. Name of department. Description of function';
COMMENT ON COLUMN table1.c3 IS 'Department Location. City where the department is located';
-- create the EMP table as TABLE2
CREATE TABLE TABLE2(
c1 NUMBER(4) not null primary key,
c2 VARCHAR2(50) not null,
c3 VARCHAR2(50) not null,
c4 NUMBER(4),
c5 DATE,
c6 NUMBER(10,2),
c7 NUMBER(10,2),
c8 NUMBER(2) not null);
COMMENT ON TABLE table2 IS 'Employee table. Contains details of each Employee. Employees';
COMMENT ON COLUMN table2.c1 IS 'Employee Number. Primary Key. Unique. How each employee is idendifed';
COMMENT ON COLUMN table2.c1 IS 'Employee Name. Name of each Employee';
COMMENT ON COLUMN table2.c3 IS 'Employee Job Title. Job Role. Current Position';
COMMENT ON COLUMN table2.c4 IS 'Manager for Employee. Manager Responsible. Who the Employee reports to';
COMMENT ON COLUMN table2.c5 IS 'Hire Date. Date the employee started in role. Commencement Date';
COMMENT ON COLUMN table2.c6 IS 'Salary. How much the employee is paid each month. Dollars';
COMMENT ON COLUMN table2.c7 IS 'Commission. How much the employee can earn each month in commission. This is extra on top of salary';
COMMENT ON COLUMN table2.c8 IS 'Department Number. Foreign Key. Join to Department Table';Annotations is a way of adding additional metadata for a database object. The Annotation is in the form of a <key, value>, which are both freeform text. The database object can have multiple Annotations.
ANNOTATIONS ([ADD|DROP] annotation_name [ annotation_value ] [ , annotation_name [ annotation_value ]... )For example, using Table 1 from about, which represents DEPT, we could add the following:
CREATE TABLE TABLE1(
...
c3 VARCHAR2(50) not null)
annotations (display 'departments');We can also add annotations are column level.
CREATE TABLE TABLE1(
c1 NUMBER(2) not null primary key ANNOTATION(key 'Department Number'),
c2 VARCHAR2(50) not null ANNOTATION(display 'Department Name. Name of department. Description of function'),
c3 VARCHAR2(50) not null) ANNOTATION(display 'Department Location. City where the department is located');At some point, only the Annotations will be passed to the LLMs, so in the meantime, you’ll need to consider the addition of Comments and Annotations.
Annotations have their own data dictionary views called USER_ANNOTATIONS, USER_ANNOTATIONS_USAGE.
Some care is needed to ensure consistency of Annotation definitions used across all database objects.
SQL History Monitoring
New in Oracle 23ai is a feature to allow tracking and monitoring the last 50 queries per session. Previously, we had other features and tools for doing this, but with SQL History we have some of this information in one location. SQL History will not replace what you’ve been using previously, but is just another tool to assist you in your monitoring and diagnostics. Each user can access their own current session history, while SQL and DBAs can view the history of all current user sessions.
If you want to use it, a user with ALTER SYTEM privilege must first change the initialisation parameter SQL_HISTORY_ENABLED instance wide to TRUE in the required PDB. The default is FALSE.
To see if it is enabled.
SELECT name,
value,
default_value,
isdefault
FROM v$parameter
WHERE name like 'sql_hist%';
NAME VALUE DEFAULT_VALUE ISDEFAULT
-------------------- ---------- ------------------------- ----------
sql_history_enabled FALSE FALSE TRUEIn the above example, the parameter is not enabled.
To enable the parameter, you need to do so using a user with SYSTEM level privileges. Use the ALTER SYSTEM privilege to enable it. For example,
alter system set sql_history_enabled=true scope=both;When you connect back to your working/developer schema you should be able to see the parameter has been enabled.
connect student/Password!@DB23
show parameter sql_history_enabled
NAME TYPE VALUE
------------------------------- ----------- ------------------------------
sql_history_enabled boolean TRUEA simple test to see it is working is to query the DUAL table
SELECT sysdate;
SYSDATE
---------
11-JUL-25
1 row selected.
SQL> select XYZ from dual;
select XYZ from dual
*
ERROR at line 1:
ORA-00904: "XYZ": invalid identifier
Help: https://docs.oracle.com/error-help/db/ora-00904/When we query the V$SQL_HISTORY view we get.
SELECT sql_text,
error_number
FROM v$sql_history;
SQL_TEXT ERROR_NUMBER
------------------------------------------------------- ------------
SELECT DECODE(USER, 'XS$NULL', XS_SYS_CONTEXT('XS$SESS 0
SELECT sysdate 0
SELECT xxx FROM dual 904[I’ve truncated the SQL_TEXT above to fit on one line. By default, only the first 100 characters of the query are visible in this view]
This V$SQL_HISTORY view becomes a little bit more interesting when you look at some of the other columns that are available. in addition to the basic information for each statement like CON_ID, SID, SESSION_SERIAL#, SQL_ID and PLAN_HASH_VALUE, there are also lots of execution statistics like ELAPSED_TIME, CPU_TIME, BUFFER_GETS or PHYSICAL_READ_REQUESTS, etc. which will be of most interest.
The full list of columns is.
DESC v$sql_history
Name Null? Type
------------------------------------------ -------- -----------------------
KEY NUMBER
SQL_ID VARCHAR2(13)
ELAPSED_TIME NUMBER
CPU_TIME NUMBER
BUFFER_GETS NUMBER
IO_INTERCONNECT_BYTES NUMBER
PHYSICAL_READ_REQUESTS NUMBER
PHYSICAL_READ_BYTES NUMBER
PHYSICAL_WRITE_REQUESTS NUMBER
PHYSICAL_WRITE_BYTES NUMBER
PLSQL_EXEC_TIME NUMBER
JAVA_EXEC_TIME NUMBER
CLUSTER_WAIT_TIME NUMBER
CONCURRENCY_WAIT_TIME NUMBER
APPLICATION_WAIT_TIME NUMBER
USER_IO_WAIT_TIME NUMBER
IO_CELL_UNCOMPRESSED_BYTES NUMBER
IO_CELL_OFFLOAD_ELIGIBLE_BYTES NUMBER
SQL_TEXT VARCHAR2(100)
PLAN_HASH_VALUE NUMBER
SQL_EXEC_ID NUMBER
SQL_EXEC_START DATE
LAST_ACTIVE_TIME DATE
SESSION_USER# NUMBER
CURRENT_USER# NUMBER
CHILD_NUMBER NUMBER
SID NUMBER
SESSION_SERIAL# NUMBER
MODULE_HASH NUMBER
ACTION_HASH NUMBER
SERVICE_HASH NUMBER
IS_FULL_SQLTEXT VARCHAR2(1)
ERROR_SIGNALLED VARCHAR2(1)
ERROR_NUMBER NUMBER
ERROR_FACILITY VARCHAR2(4)
STATEMENT_TYPE VARCHAR2(5)
IS_PARALLEL VARCHAR2(1)
CON_ID NUMBEROCI Stored Video Analysis
OCI Video Analysis is a part of the OCI Vision service, designed to process stored videos and identify labels, objects, texts, and faces within each frame. It can analyze the entire video or every frame in the video using pre-trained or custom models. The feature provides information about the detected labels, objects, texts, and faces, and provides the time at which they’re detected. With a timeline bar, you can directly look for a label or object and navigate to the exact timestamp in the video where a particular label or object is found.
To use this feature, you’ll need to upload your video to an OCI Bucket. Here is an example of a video stored in a bucket called OCI-Vision-Video-Demos.
You might need to allow Pre-Authenticated Requests for this bucket. If you don’t do this, you will be prompted by the tool to allow this.
Next, go to the OCI Vision page. You’ll find the Video Analysis tool at the bottom of the menu on the left-hand side of the page.
You can check out the demo videos, or load your own video from the Local File system of you computer, or use a file from your OCI Storage. If you select a video from the Local File system, the video will be loaded in the object storage before it is processed.
For this example, I’m going to use the video I uploaded earlier called Trinity-Student.mp4. Copy the link to this file from the file information in object storage.
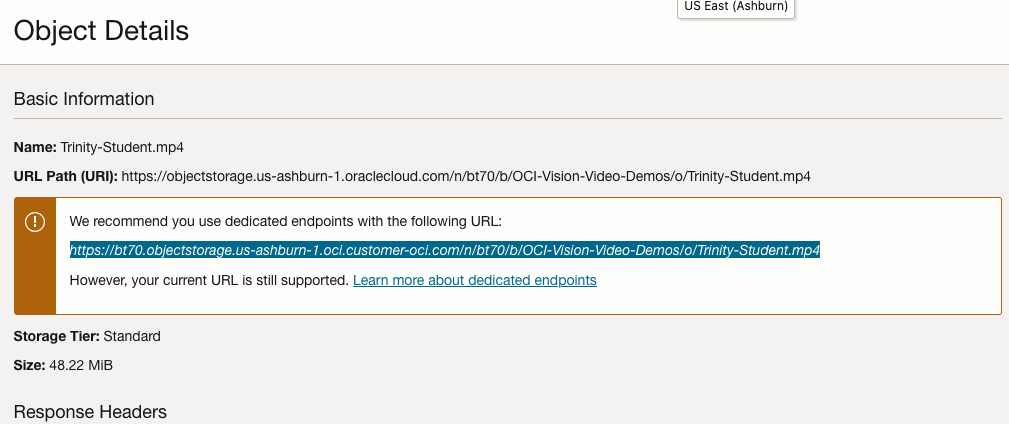
On the Video Analysis page, select Object Storage and paste the link to the file into the URL field. Then click Analyze button. It is at this point that you might get asked to Generate a PAR URL. Do this and then proceed.
While the video is being parsed, it will appear on the webpage and will start playing. When the video has been Analyzed the details will be displayed below the video. The Analysis will consist of Label Detection, Object Detection, Text Dection and Face Detection.
By clicking on each of these, you’ll see what has been detected, and by clicking on each of these, you’ll be able to see where in the video they were detected. For example, where a Chair was detected.
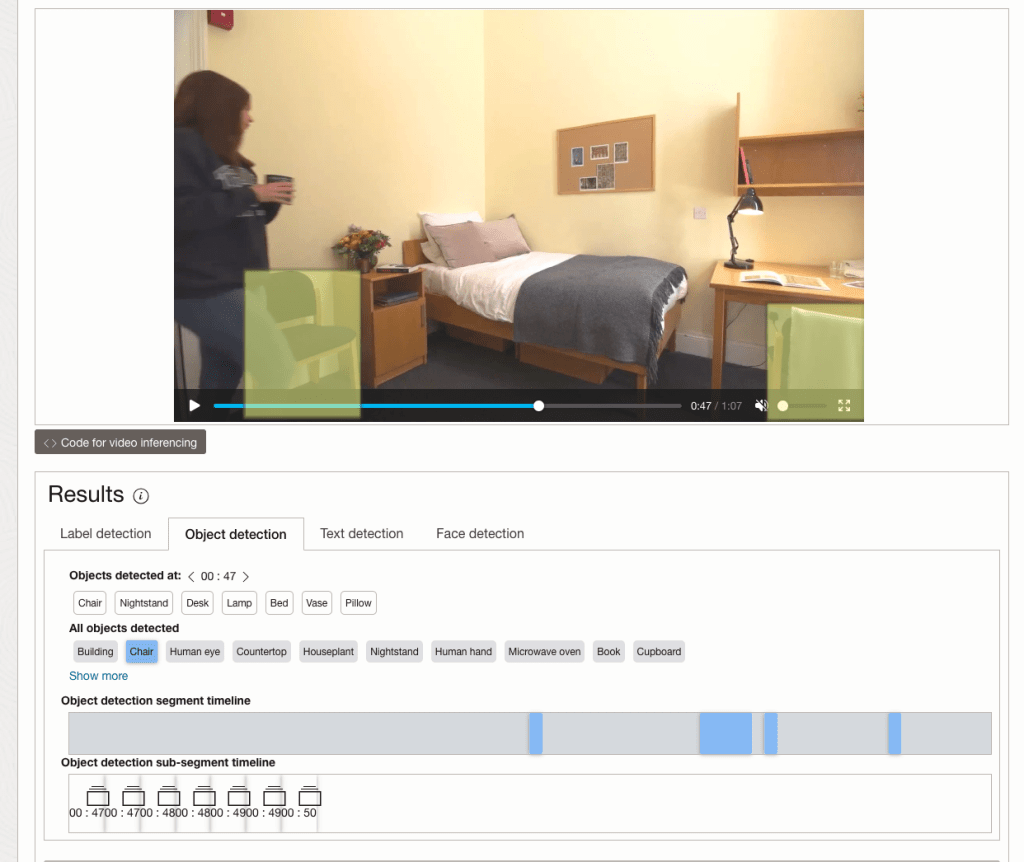
You can also inspect the JSON file containing all the details of various objects detected in the video and the location in the videos they can be found.
This JSON response file is also saved to Object Storage in the same directory, or a sub-directory, where the video is located.
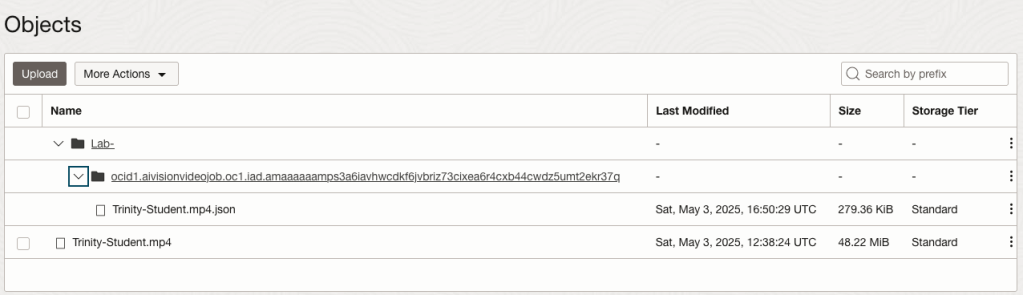
OCI Text to Speech example
In this post, I’ll walk through the steps to get a very simple example of Text-to-Speech working. This example builds upon my previous posts on OCI Language, OCI Speech and others, so make sure you check out those posts.
The first thing you need to be aware of, and to check, before you proceed, is whether the Text-to-Speech is available in your region. At the time of writing, this feature was only available in Phoenix, which is one of the cloud regions I have access to. There are plans to roll it out to other regions, but I’m not aware of the timeline for this. Although you might see Speech listed on your AI menu in OCI, that does not guarantee the Text-to-Speech feature is available. What it does mean is the text trans scribing feature is available.
So if Text-to-Speech is available in your region, the following will get you up and running.
The first thing you need to do is read in the Config file from the OS.
#initial setup, read Config file, create OCI Client
import oci
from oci.config import from_file
##########
from oci_ai_speech_realtime import RealtimeSpeechClient, RealtimeSpeechClientListener
from oci.ai_speech.models import RealtimeParameters
##########
CONFIG_PROFILE = "DEFAULT"
config = oci.config.from_file('~/.oci/config', profile_name=CONFIG_PROFILE)
###
ai_speech_client = ai_speech_client = oci.ai_speech.AIServiceSpeechClient(config)
###
print(config)
### Update region to point to Phoenix
config.update({'region':'us-phoenix-1'})A simple little test to see if the Text-to-Speech feature is enabled for your region is to display the available list of voices.
list_voices_response = ai_speech_client.list_voices(
compartment_id=COMPARTMENT_ID,
display_name="Text-to-Speech")
# opc_request_id="1GD0CV5QIIS1RFPFIOLF<unique_ID>")
# Get the data from response
print(list_voices_response.data)This produces a long json object with many characteristics of the available voices. A simpler listing gives the names and gender)
for i in range(len(list_voices_response.data.items)):
print(list_voices_response.data.items[i].display_name + ' [' + list_voices_response.data.items[i].gender + ']\t' + list_voices_response.data.items[i].language_description )
------
Brian [MALE] English (United States)
Annabelle [FEMALE] English (United States)
Bob [MALE] English (United States)
Stacy [FEMALE] English (United States)
Phil [MALE] English (United States)
Cindy [FEMALE] English (United States)
Brad [MALE] English (United States)
Richard [MALE] English (United States)Now lets setup a Text-to-Speech example using the simple text, Hello. My name is Brendan and this is an example of using Oracle OCI Speech service. First lets define a function to save the audio to a file.
def save_audi_response(data):
with open(filename, 'wb') as f:
for b in data.iter_content():
f.write(b)
f.close()We can now establish a connection, define the text, call the OCI Speech function to create the audio, and then to save the audio file.
import IPython.display as ipd
# Initialize service client with default config file
ai_speech_client = oci.ai_speech.AIServiceSpeechClient(config)
TEXT_DEMO = "Hello. My name is Brendan and this is an example of using Oracle OCI Speech service"
#speech_response = ai_speech_client.synthesize_speech(compartment_id=COMPARTMENT_ID)
speech_response = ai_speech_client.synthesize_speech(
synthesize_speech_details=oci.ai_speech.models.SynthesizeSpeechDetails(
text=TEXT_DEMO,
is_stream_enabled=True,
compartment_id=COMPARTMENT_ID,
configuration=oci.ai_speech.models.TtsOracleConfiguration(
model_family="ORACLE",
model_details=oci.ai_speech.models.TtsOracleTts2NaturalModelDetails(
model_name="TTS_2_NATURAL",
voice_id="Annabelle"),
speech_settings=oci.ai_speech.models.TtsOracleSpeechSettings(
text_type="SSML",
sample_rate_in_hz=18288,
output_format="MP3",
speech_mark_types=["WORD"])),
audio_config=oci.ai_speech.models.TtsBaseAudioConfig(config_type="BASE_AUDIO_CONFIG") #, save_path='I'm not sure what this should be')
) )
# Get the data from response
#print(speech_response.data)
save_audi_response(speech_response.data)Translating Text using OCI AI Services
I’ve written several blog posts on using various features and functions of the OCI AI Services, and the most recent of these have been about some of the Language features. In this blog post, I’ll show you how to use the OCI Language Translation service.
As with the previous posts, there is some initial configuration and setup for your computer to access the OCI cloud services. Check out my previous posts on this. The following examples assume you have that configuration setup.
The OCI Translation service can translate text into over 30 different languages, with more being added over time.
There are 3 things needed to use the Translation Service. The Text to be translated, what language that text is in and what language you’d like the text translated into. Sounds simple. Well, it kind of is, but some care is needed to ensure it all works smoothly.
Let’s start with the basic setup of importing libraries, reading the config file and initialising the OCI AI Client.
import oci
from oci.config import from_file
#Read in config file - this is needed for connecting to the OCI AI Services
#COMPARTMENT_ID = "ocid1.tenancy.oc1..aaaaaaaaop3yssfqnytz5uhc353cmel22duc4xn2lnxdr4f4azmi2fqga4qa"
CONFIG_PROFILE = "DEFAULT"
config = oci.config.from_file('~/.oci/config', profile_name=CONFIG_PROFILE)
###
ai_language_client = oci.ai_language.AIServiceLanguageClient(config)
Next, we can define what text we want to translate and what Language we want to translate the text into. In this case, I want to translate the text into French and to do so, we need to use the language abbreviation.
text_to_trans = "Hello. My name is Brendan and this is an example of using Oracle OCI Language translation service"
print(text_to_trans)
target_lang = "fr"Next, we need to prepare the text and then send it to the translation service. Then, print the returned object
t_doc = oci.ai_language.models.TextDocument(key="Demo", text=text_to_trans, language_code="en")
trans_response = ai_language_client.batch_language_translation(oci.ai_language.models.BatchLanguageTranslationDetails(documents=[t_doc], target_language_code=target_lang))
print(trans_response.data)The returned translated object is the following.
{
"documents": [
{
"key": "Demo",
"source_language_code": "en",
"target_language_code": "fr",
"translated_text": "Bonjour. Je m'appelle Brendan et voici un exemple d'utilisation du service de traduction Oracle OCI Language"
}
],
"errors": []
}We can automate this process a little to automatically detect the input language. For example:
source_lang = ai_language_client.detect_dominant_language(oci.ai_language.models.DetectLanguageSentimentsDetails(text=text_to_trans))
t_doc = oci.ai_language.models.TextDocument(key="Demo", text=text_to_trans, language_code=source_lang.data.languages[0].code)
trans_response = ai_language_client.batch_language_translation(oci.ai_language.models.BatchLanguageTranslationDetails(documents=[t_doc], target_language_code=target_lang))
print(trans_response.data)
And we can also automate the translation into multiple different langues.
text_to_trans = "Hello. My name is Brendan and this is an example of using Oracle OCI Language translation service"
print(text_to_trans)
#target_lang = "fr"
target_lang = {"fr":"French", "nl":"Dutch", "de":"German", "it":"Italian", "ja":"Japaneese", "ko":"Korean", "pl":"polish"}
for lang_key, lang_name in target_lang.items():
t_doc = oci.ai_language.models.TextDocument(key="Demo", text=text_to_trans, language_code="en")
trans_response = ai_language_client.batch_language_translation(oci.ai_language.models.BatchLanguageTranslationDetails(documents=[t_doc], target_language_code=lang_key))
####
print(' [' + lang_name + '] ' + trans_response.data.documents[0].translated_text)Hello. My name is Brendan and this is an example of using Oracle OCI Language translation service
[French] Bonjour. Je m'appelle Brendan et voici un exemple d'utilisation du service de traduction Oracle OCI Language
[Dutch] Hallo. Mijn naam is Brendan en dit is een voorbeeld van het gebruik van de Oracle OCI Language vertaalservice.
[German] Hallo. Mein Name ist Brendan und dies ist ein Beispiel für die Verwendung des Oracle OCI Language-Übersetzungsservice
[Italian] Ciao. Il mio nome è Brendan e questo è un esempio di utilizzo del servizio di traduzione di Oracle OCI Language
[Japaneese] こんにちは。私の名前はBrendanで、これはOracle OCI Language翻訳サービスの使用例です
[Korean] 안녕하세요. 내 이름은 브렌단이며 Oracle OCI 언어 번역 서비스를 사용하는 예입니다.
[polish] Dzień dobry. Nazywam się Brendan i jest to przykład korzystania z usługi tłumaczeniowej OCI LanguageUnlock Text Analytics with Oracle OCI Python – Part 2
This is my second post on using Oracle OCI Language service to perform Text Analytics. These include Language Detection, Text Classification, Sentiment Analysis, Key Phrase Extraction, Named Entity Recognition, Private Data detection and masking, and Healthcare NLP.
In my Previous post (Part 1), I covered examples on Language Detection, Text Classification and Sentiment Analysis.
In this post (Part 2), I’ll cover:
- Key Phrase
- Named Entity Recognition
- Detect private information and marking
Make sure you check out Part 1 for details on setting up the client and establishing a connection. These details are omitted in the examples below.
Key Phrase Extraction
With Key Phrase Extraction, it aims to identify the key works and/or phrases from the text. The keywords/phrases are selected based on what are the main topics in the text along with the confidence score. The text is parsed to extra the words/phrase that are important in the text. This can aid with identifying the key aspects of the document without having to read it. Care is needed as these words/phrases do not represent the meaning implied in the text.
Using some of the same texts used in Part-1, let’s see what gets generated for the text about a Hotel experience.
t_doc = oci.ai_language.models.TextDocument(
key="Demo",
text="This hotel is a bad place, I would strongly advise against going there. There was one helpful member of staff",
language_code="en")
key_phrase = ai_language_client.batch_detect_language_key_phrases((oci.ai_language.models.BatchDetectLanguageKeyPhrasesDetails(documents=[t_doc])))
print(key_phrase.data)
print('==========')
for i in range(len(key_phrase.data.documents)):
for j in range(len(key_phrase.data.documents[i].key_phrases)):
print("phrase: ", key_phrase.data.documents[i].key_phrases[j].text +' [' + str(key_phrase.data.documents[i].key_phrases[j].score) + ']'){
"documents": [
{
"key": "Demo",
"key_phrases": [
{
"score": 0.9998106383818767,
"text": "bad place"
},
{
"score": 0.9998106383818767,
"text": "one helpful member"
},
{
"score": 0.9944029848214838,
"text": "staff"
},
{
"score": 0.9849306609397931,
"text": "hotel"
}
],
"language_code": "en"
}
],
"errors": []
}
==========
phrase: bad place [0.9998106383818767]
phrase: one helpful member [0.9998106383818767]
phrase: staff [0.9944029848214838]
phrase: hotel [0.9849306609397931]The output from the Key Phrase Extraction is presented into two formats about. The first is the JSON object returned from the function, containing the phrases and their confidence score. The second (below the ==========) is a formatted version of the same JSON object but parsed to extract and present the data in a more compact manner.
The next piece of text to be examined is taken from an article on the F1 website about a change of divers.
text_f1 = "Red Bull decided to take swift action after Liam Lawsons difficult start to the 2025 campaign, demoting him to Racing Bulls and promoting Yuki Tsunoda to the senior team alongside reigning world champion Max Verstappen. F1 Correspondent Lawrence Barretto explains why… Sergio Perez had endured a painful campaign that saw him finish a distant eighth in the Drivers Championship for Red Bull last season – while team mate Verstappen won a fourth successive title – and after sticking by him all season, the team opted to end his deal early after Abu Dhabi finale."
t_doc = oci.ai_language.models.TextDocument(
key="Demo",
text=text_f1,
language_code="en")
key_phrase = ai_language_client.batch_detect_language_key_phrases(oci.ai_language.models.BatchDetectLanguageKeyPhrasesDetails(documents=[t_doc]))
print(key_phrase.data)
print('==========')
for i in range(len(key_phrase.data.documents)):
for j in range(len(key_phrase.data.documents[i].key_phrases)):
print("phrase: ", key_phrase.data.documents[i].key_phrases[j].text +' [' + str(key_phrase.data.documents[i].key_phrases[j].score) + ']')I won’t include all the output and the following shows the key phrases in the compact format
phrase: red bull [0.9991468440416812]
phrase: swift action [0.9991468440416812]
phrase: liam lawsons difficult start [0.9991468440416812]
phrase: 2025 campaign [0.9991468440416812]
phrase: racing bulls [0.9991468440416812]
phrase: promoting yuki tsunoda [0.9991468440416812]
phrase: senior team [0.9991468440416812]
phrase: sergio perez [0.9991468440416812]
phrase: painful campaign [0.9991468440416812]
phrase: drivers championship [0.9991468440416812]
phrase: red bull last season [0.9991468440416812]
phrase: team mate verstappen [0.9991468440416812]
phrase: fourth successive title [0.9991468440416812]
phrase: all season [0.9991468440416812]
phrase: abu dhabi finale [0.9991468440416812]
phrase: team [0.9420016064526977]While some aspects of this is interesting, care is needed to not overly rely upon it. It really depends on the usecase.
Named Entity Recognition
For Named Entity Recognition is a natural language process for finding particular types of entities listed as words or phrases in the text. The named entities are a defined list of items. For OCI Language there is a list available here. Some named entities have a sub entities. The return JSON object from the function has a format like the following.
{
"documents": [
{
"entities": [
{
"length": 5,
"offset": 5,
"score": 0.969588577747345,
"sub_type": "FACILITY",
"text": "hotel",
"type": "LOCATION"
},
{
"length": 27,
"offset": 82,
"score": 0.897526216506958,
"sub_type": null,
"text": "one helpful member of staff",
"type": "QUANTITY"
}
],
"key": "Demo",
"language_code": "en"
}
],
"errors": []
}For each named entity discovered the returned object will contain the Text identifed, the Entity Type, the Entity Subtype, Confidence Score, offset and length.
Using the text samples used previous, let’s see what gets produced. The first example is for the hotel review.
t_doc = oci.ai_language.models.TextDocument(
key="Demo",
text="This hotel is a bad place, I would strongly advise against going there. There was one helpful member of staff",
language_code="en")
named_entities = ai_language_client.batch_detect_language_entities(
batch_detect_language_entities_details=oci.ai_language.models.BatchDetectLanguageEntitiesDetails(documents=[t_doc]))
for i in range(len(named_entities.data.documents)):
for j in range(len(named_entities.data.documents[i].entities)):
print("Text: ", named_entities.data.documents[i].entities[j].text, ' [' + named_entities.data.documents[i].entities[j].type + ']'
+ '[' + str(named_entities.data.documents[i].entities[j].sub_type) + ']' + '{offset:'
+ str(named_entities.data.documents[i].entities[j].offset) + '}')Text: hotel [LOCATION][FACILITY]{offset:5}
Text: one helpful member of staff [QUANTITY][None]{offset:82}The last two lines above are the formatted output of the JSON object. It contains two named entities. The first one is for the text “hotel” and it has a Entity Type of Location, and a Sub Entitity Type of Location. The second named entity is for a long piece of string and for this it has a Entity Type of Quantity, but has no Sub Entity Type.
Now let’s see what is creates for the F1 text. (the text has been given above and the code is very similar/same as above).
Text: Red Bull [ORGANIZATION][None]{offset:0}
Text: swift [ORGANIZATION][None]{offset:25}
Text: Liam Lawsons [PERSON][None]{offset:44}
Text: 2025 [DATETIME][DATE]{offset:80}
Text: Yuki Tsunoda [PERSON][None]{offset:138}
Text: senior [QUANTITY][AGE]{offset:158}
Text: Max Verstappen [PERSON][None]{offset:204}
Text: F1 [ORGANIZATION][None]{offset:220}
Text: Lawrence Barretto [PERSON][None]{offset:237}
Text: Sergio Perez [PERSON][None]{offset:269}
Text: campaign [EVENT][None]{offset:304}
Text: eighth in the [QUANTITY][None]{offset:343}
Text: Drivers Championship [EVENT][None]{offset:357}
Text: Red Bull [ORGANIZATION][None]{offset:382}
Text: Verstappen [PERSON][None]{offset:421}
Text: fourth successive title [QUANTITY][None]{offset:438}
Text: Abu Dhabi [LOCATION][GPE]{offset:545}Detect Private Information and Marking
The ability to perform data masking has been available in SQL for a long time. There are lots of scenarios where masking is needed and you are not using a Database or not at that particular time.
With Detect Private Information or Personal Identifiable Information the OCI AI function search for data that is personal and gives you options on how to present this back to the users. Examples of the types of data or Entity Types it will detect include Person, Adddress, Age, SSN, Passport, Phone Numbers, Bank Accounts, IP Address, Cookie details, Private and Public keys, various OCI related information, etc. The list goes on. Check out the documentation for more details on these. Unfortunately the documentation for how the Python API works is very limited.
The examples below illustrate some of the basic options. But there is lots more you can do with this feature like defining you own rules.
For these examples, I’m going to use the following text which I’ve assigned to a variable called text_demo.
Hi Martin. Thanks for taking my call on 1/04/2025. Here are the details you requested. My Bank Account Number is 1234-5678-9876-5432 and my Bank Branch is Main Street, Dublin. My Date of Birth is 29/02/1993 and I’ve been living at my current address for 15 years. Can you also update my email address to brendan.tierney@email.com. If toy have any problems with this you can contact me on +353-1-493-1111. Thanks for your help. Brendan.
m_mode = {"ALL":{"mode":'MASK'}}
t_doc = oci.ai_language.models.TextDocument(key="Demo", text=text_demo,language_code="en")
pii_entities = ai_language_client.batch_detect_language_pii_entities(oci.ai_language.models.BatchDetectLanguagePiiEntitiesDetails(documents=[t_doc], masking=m_mode))
print(text_demo)
print('--------------------------------------------------------------------------------')
print(pii_entities.data.documents[0].masked_text)
print('--------------------------------------------------------------------------------')
for i in range(len(pii_entities.data.documents)):
for j in range(len(pii_entities.data.documents[i].entities)):
print("phrase: ", pii_entities.data.documents[i].entities[j].text +' [' + str(pii_entities.data.documents[i].entities[j].type) + ']')
Hi Martin. Thanks for taking my call on 1/04/2025. Here are the details you requested. My Bank Account Number is 1234-5678-9876-5432 and my Bank Branch is Main Street, Dublin. My Date of Birth is 29/02/1993 and I've been living at my current address for 15 years. Can you also update my email address to brendan.tierney@email.com. If toy have any problems with this you can contact me on +353-1-493-1111. Thanks for your help. Brendan.
--------------------------------------------------------------------------------
Hi ******. Thanks for taking my call on *********. Here are the details you requested. My Bank Account Number is ******************* and my Bank Branch is Main Street, Dublin. My Date of Birth is ********** and I've been living at my current address for ********. Can you also update my email address to *************************. If toy have any problems with this you can contact me on ***************. Thanks for your help. *******.
--------------------------------------------------------------------------------
phrase: Martin [PERSON]
phrase: 1/04/2025 [DATE_TIME]
phrase: 1234-5678-9876-5432 [CREDIT_DEBIT_NUMBER]
phrase: 29/02/1993 [DATE_TIME]
phrase: 15 years [DATE_TIME]
phrase: brendan.tierney@email.com [EMAIL]
phrase: +353-1-493-1111 [TELEPHONE_NUMBER]
phrase: Brendan [PERSON]The above this the basic level of masking.
A second option is to use the REMOVE mask. For this, change the mask format to the following.
m_mode = {"ALL":{'mode':'REMOVE'}} For this option the indentified information is removed from the text.
Hi . Thanks for taking my call on . Here are the details you requested. My Bank Account Number is and my Bank Branch is Main Street, Dublin. My Date of Birth is and I've been living at my current address for . Can you also update my email address to . If toy have any problems with this you can contact me on . Thanks for your help. .
--------------------------------------------------------------------------------
phrase: Martin [PERSON]
phrase: 1/04/2025 [DATE_TIME]
phrase: 1234-5678-9876-5432 [CREDIT_DEBIT_NUMBER]
phrase: 29/02/1993 [DATE_TIME]
phrase: 15 years [DATE_TIME]
phrase: brendan.tierney@email.com [EMAIL]
phrase: +353-1-493-1111 [TELEPHONE_NUMBER]
phrase: Brendan [PERSON]For the REPLACE option we have.
m_mode = {"ALL":{'mode':'REPLACE'}} Which gives us the following, where we can see the key information is removed and replace with the name of Entity Type.
Hi <PERSON>. Thanks for taking my call on <DATE_TIME>. Here are the details you requested. My Bank Account Number is <CREDIT_DEBIT_NUMBER> and my Bank Branch is Main Street, Dublin. My Date of Birth is <DATE_TIME> and I've been living at my current address for <DATE_TIME>. Can you also update my email address to <EMAIL>. If toy have any problems with this you can contact me on <TELEPHONE_NUMBER>. Thanks for your help. <PERSON>.
--------------------------------------------------------------------------------
phrase: Martin [PERSON]
phrase: 1/04/2025 [DATE_TIME]
phrase: 1234-5678-9876-5432 [CREDIT_DEBIT_NUMBER]
phrase: 29/02/1993 [DATE_TIME]
phrase: 15 years [DATE_TIME]
phrase: brendan.tierney@email.com [EMAIL]
phrase: +353-1-493-1111 [TELEPHONE_NUMBER]
phrase: Brendan [PERSON]We can also change the character used for the masking. In this example we change the masking character to + symbol.
m_mode = {"ALL":{'mode':'MASK','maskingCharacter':'+'}} Hi ++++++. Thanks for taking my call on +++++++++. Here are the details you requested. My Bank Account Number is +++++++++++++++++++ and my Bank Branch is Main Street, Dublin. My Date of Birth is ++++++++++ and I've been living at my current address for ++++++++. Can you also update my email address to +++++++++++++++++++++++++. If toy have any problems with this you can contact me on +++++++++++++++. Thanks for your help. +++++++.I mentioned at the start of this section there was lots of options available to you, including defining your own rules, using regular expressions, etc. Let me know if you’re interested in exploring some of these and I can share a few more examples.
- ← Previous
- 1
- 2
- 3
- …
- 57
- Next →


You must be logged in to post a comment.